PyNutil#
a Python library for brain-wide quantification and spatial analysis of features in serial section images from the brain.
Installation, supported formats and key concepts.
Use PyNutil via the graphical user interface.
A gallery of examples using PyNutil.
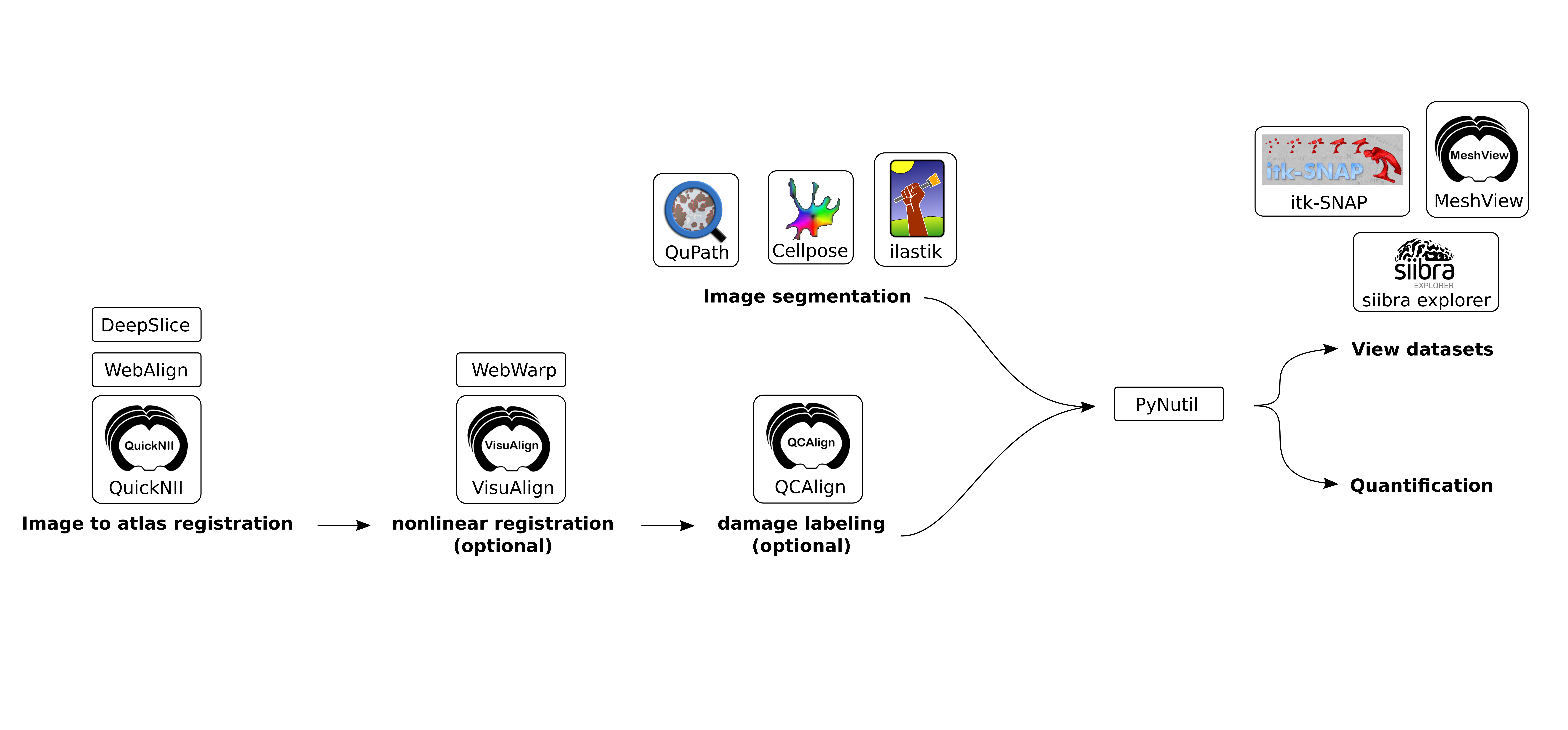
Overview#
PyNutil is able to integrate outputs from various atlas registration software and image segmentation software in order to produce atlas based quantifications, 3D point clouds, and 3D heatmaps of brain derived data.
PyNutil aims to replicate and expand the Quantifier feature of the Nutil software (RRID: SCR_017183).
Warning
PyNutil is still under development and the API is subject to change.
The documentation below brings together installation notes, the basic workflow, practical examples, and the generated API reference.
Contents:
Highlights#
Use BrainGlobe atlases or custom atlas volumes in
.nrrdformat.Read registration data from QuickNII, VisuAlign, DeepSlice, or BrainGlobe registration outputs.
Quantify binary segmentations or intensity images against atlas regions.
Export point clouds for MeshView and interpolated NIfTI volumes for siibra explorer or ITK-SNAP.
For more background on the QUINT workflow, see QUINT workflow documentation.