Visualising PyNutil volumes in brainrender#
Render a PyNutil interpolated expression volume as a semi-transparent 3D heatmap in brainrender, with no intermediate files written to disk.
The example below uses a single calbindin-1 ISH section series registered with section-to-volume alignment only.
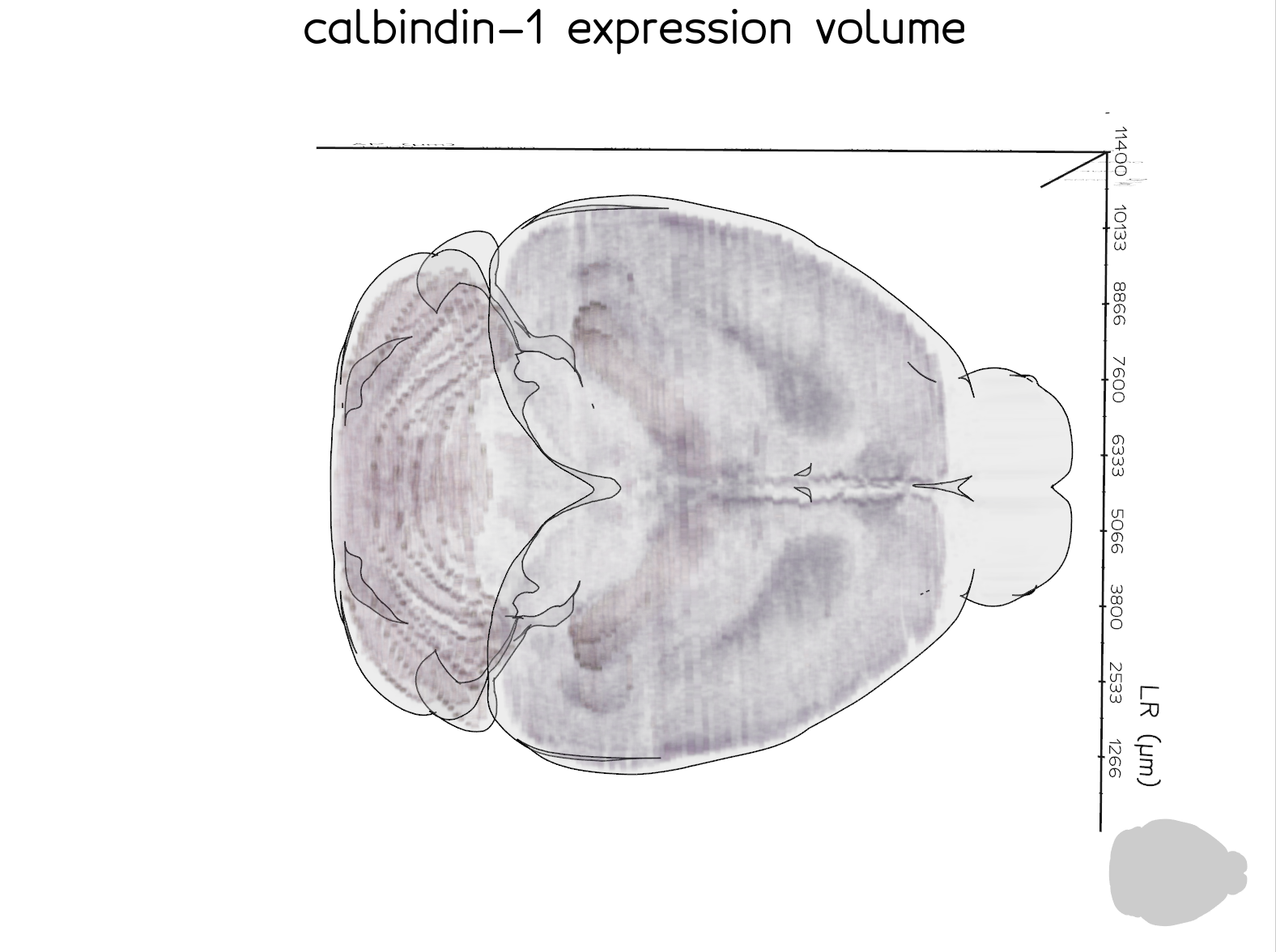
A more fully-powered example is the Rorb dataset below, which combines 26 section series and nonlinear (ANTS) registration to produce a smoother, more complete volume. This dataset is from the study available at https://doi.org/10.64898/2026.01.20.700446.
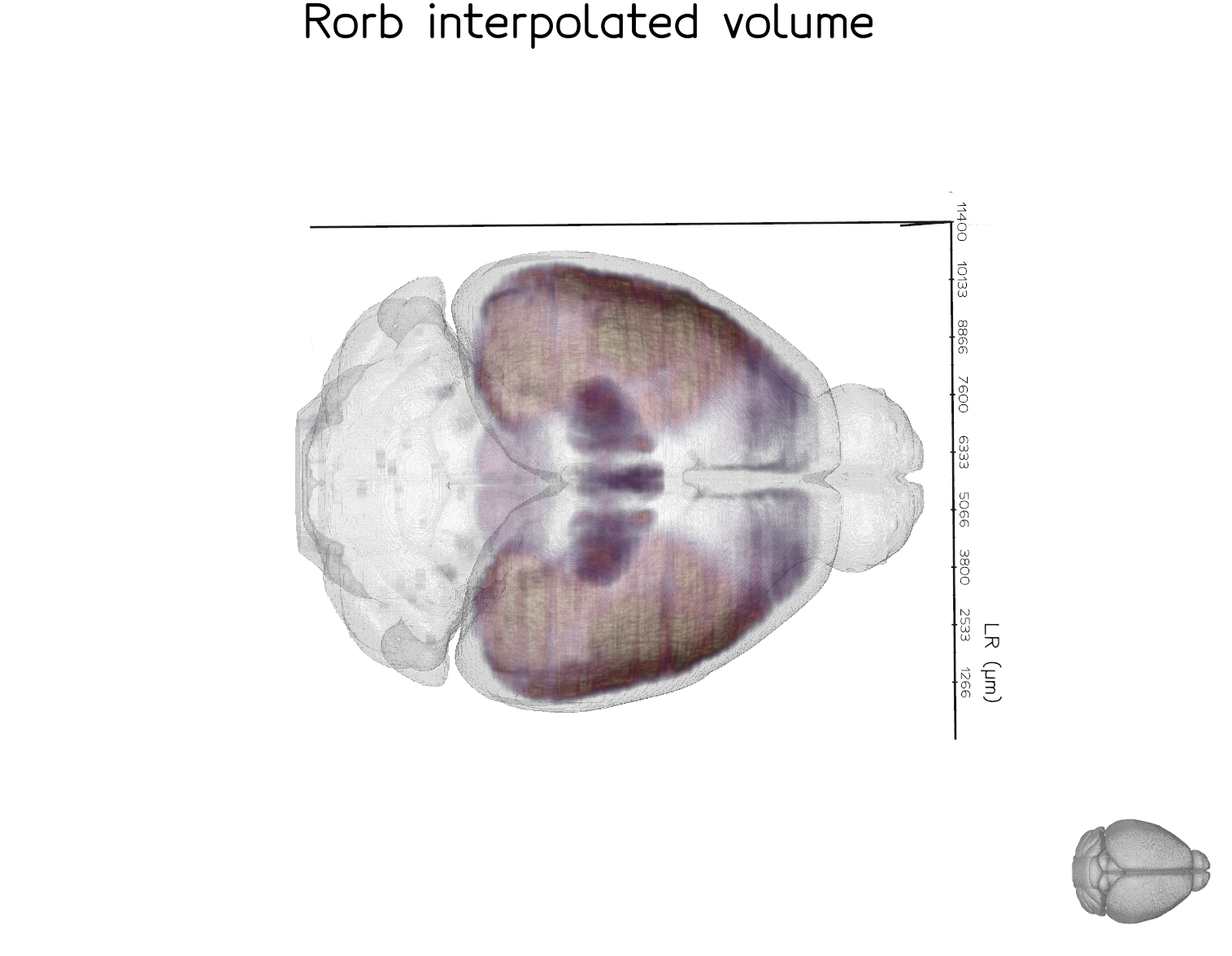
Code#
import numpy as np
from brainglobe_atlasapi import BrainGlobeAtlas
from brainrender import Scene, settings
from brainrender.actors import Volume
import PyNutil as pnt
settings.SHADER_STYLE = "default"
IMAGE_DIR = "path/to/expression_25um"
ALIGNMENT_JSON = "path/to/alignment.json"
atlas = BrainGlobeAtlas("allen_mouse_25um")
alignment = pnt.read_alignment(ALIGNMENT_JSON)
image_series = pnt.read_image_dir(IMAGE_DIR)
result = pnt.interpolate_volume(
image_series=image_series,
registration=alignment,
atlas=atlas,
value_mode="mean",
segmentation_mode=False,
intensity_channel="grayscale",
do_interpolation=True,
return_orientation="rsa", # required for brainrender axis convention
)
arr = result.value.astype(np.float32)
arr = np.nan_to_num(arr, nan=0.0)
vmax = float(arr.max())
scene = Scene(atlas_name="allen_mouse_25um", title="expression volume")
volume = Volume(
arr,
voxel_size=25,
cmap="magma",
as_surface=False,
)
volume.mesh.alpha([
(0, 0.0),
(1, 0.0),
(vmax * 0.2, 0.10),
(vmax * 0.5, 0.2),
(vmax * 0.8, 0.4),
(vmax, 0.6),
])
volume.mesh.cmap("magma", vmin=0, vmax=vmax * 0.6)
volume.mesh.alpha_unit(300)
volume.mesh.mode(0)
scene.add(volume)
scene.render()